Kidney biology / Nephrology
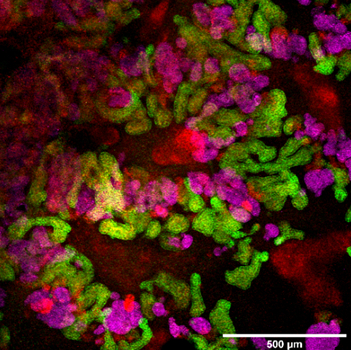
iPSCs-derived kidney organoid at day 26 expressing CDH1 (green), GATA3 (red) and NPHS1 (magenta)
Image courtesy Bachmann lab (Joana Figueiro da Silva)
________________________________________________________________
LABS |
|
__________________________________________________________________
|
|
Prof. Dr. med. Ruxandra Bachmann-Gagescu Associate Professor for Developmental Genetics Department of Molecular Life Sciences and Institute of Medical Genetics, University of Zurich Research Focus: Our research focuses on a group of human Mendelian disorders called ciliopathies, which are unified by shared genetic causes resulting in primary cilium dysfunction. Primary cilia are small non-motile organelles present on the surface of most vertebrate cells where they are involved in transduction of sensory, mechanical or chemical signals and in regulation of signaling pathways during development and cell homeostasis. Typical clinical presentations of ciliopathies include neurological involvement, retinal degeneration and renal fibrocystic disease, as illustrated by Joubert syndrome (JS), an iconic ciliopathy, which is the main focus of our research. To understand the consequences of mutations in JS-associated genes at the molecular level, we are developing human iPSC-based renal organoids using CRISPR/Cas9 genome editing, imaging and –omics approaches such as RNAsequencing and proteomics. Methods: CRISPR/Cas9 genome editing, next generation sequencing for clone selection and quality control, 2D cortical neuronal differentiation, 3D cerebral organoids, renal organoids Keywords: ciliopathies, primary cilia, Joubert syndrome, iPSCs, neurons, renal tubular cells Topics: Neuroscience, Disease Modelling, Kidney biology Publications:https://pubmed.ncbi.nlm.nih.gov/?term=ruxandra+Bachmann-Gagescu&sort=date&size=100 Website: https://www.medgen.uzh.ch/en/forschung/gagescu.html _____________________________________________________________________top |
|
|
Prof. Dr. med. Soeren Lienkamp Assistant Professor Institute of Anatomy, University of Zurich Research Focus: We are interested in embryonic renal organogenesis and the pathophysiology of various genetic renal disorders, including congenital renal anomalies, ciliopathies and cystic kidney disease. We also aim to understand how transcriptional control mechanisms influence renal tubular cell identity. To model genetic diseases, we use in vivo (Xenopus tropicalis embryos) and vitro (direct reprogramming of fibroblasts to renal-like epithelial cells) approaches in combination with CRISPR/Cas9 genome editing, light-sheet microscopy, and machine learning. We are collaborating with the group of Ruxandra Bachmann-Gagescu to differentiate iPSCs into renal organoids to understand the molecular alterations in ciliopathies and kidney malformations. Methods: direct reprogramming, CRISPR/Cas9 genome editing, Xenopus tropicalis embryology, light sheet microscopy, deep learning-based image analysis, renal organoids Keywords: kidney, ciliopathies, cystic kidney disease, Xenopus, renal tubular cells Topics: Renal organogenesis, Kidney disease modelling Publications: https://pubmed.ncbi.nlm.nih.gov/?term=lienkamp+s&sort=date Website: https://www.anatomy.uzh.ch/en/research/lienkamp.html __________________________________________________________________________top |

